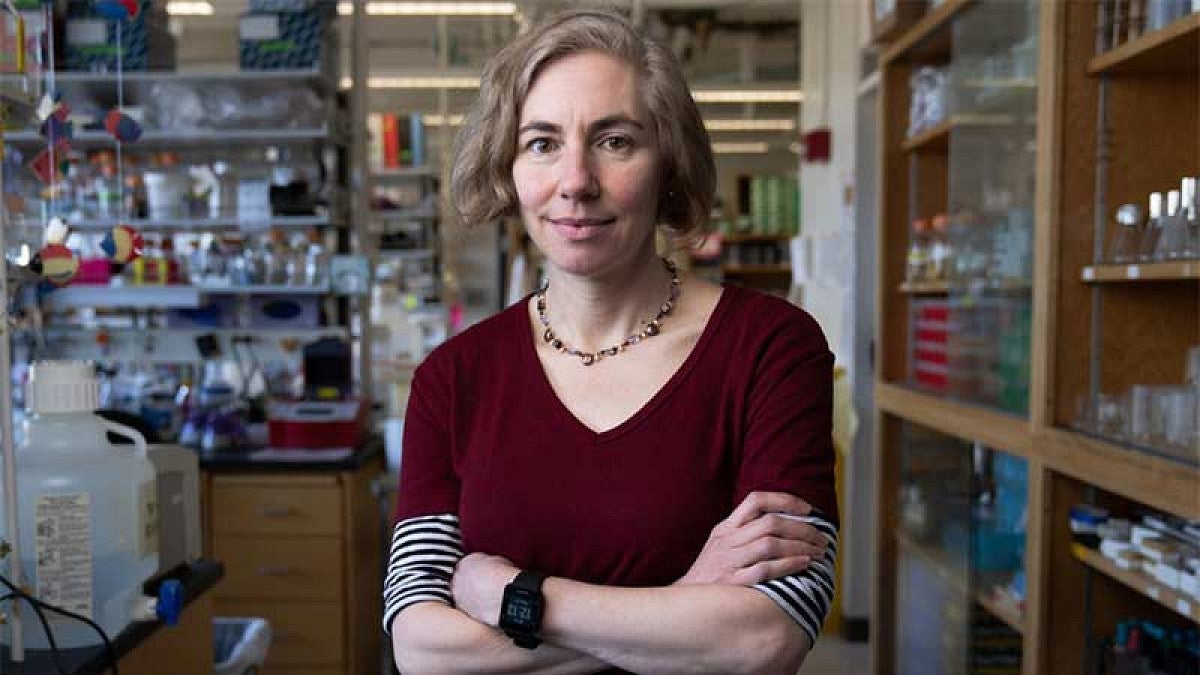
The COVID-19 (coronavirus) pandemic illustrates how all of us are interconnected by microbes. While this virus has spread disease through homes and workplaces, other microbes that circulate among humans can be benign or even beneficial. This world of people’s shared microbial communities, or “microbiomes,” is the domain of Karen Guillemin, Philip H. Knight Chair and professor of biology.
When she joined the University of Oregon’s Institute of Molecular Biology as a junior professor in 2001, Guillemin hoped to create microbiologically sterile (“germfree”) zebrafish because they would allow her to explore how individual types of bacteria affect our health, one species at a time. UO researchers already had established zebrafish, which share 70 percent of our genes, as an ideal model for studying human disease.
“I knew Oregon was the central place for genetic research with zebrafish,” Guillemin says. “I came here with the idea of using zebrafish to understand how we coexist with microbes.”
RELATED LINKS
Success came five years later, with a breakthrough 2006 publication describing how germfree zebrafish development is stunted in specific ways. Since then, germfree zebrafish have helped Guillemin discover bacterial proteins that simultaneously help bacteria compete for real estate in the zebrafish gut and affect zebrafish biology. The process of discovering these novel proteins is roughly comparable to sifting through tons of sand in a search for a single grain that is unlike the rest. These findings are pointing the way to new therapeutics for inflammation and diseases such as diabetes.
“We are realizing that the microbiome is a rich source for discovering new biomolecules,” she says. “They have enormous potential for manipulating and promoting our health.”
Guillemin traces her fascination with biology to childhood summers spent exploring tide pools in Cape Cod. Although she grew up in an academic household in Cambridge, Massachusetts, with a mathematician father and Germanist mother, she didn’t have any sense of what a career in biology might be like.
She majored in biology at Harvard, but considered switching to philosophy midway through her sophomore year because she found it far more stimulating than entry-level science courses. “I was interested in biology, but I was not excited by my classroom experiences,” she says. “The big thing that got me excited was joining a lab and learning what it was like to do experimental science, which is entirely different from sitting in class and being told things.”
By cold-calling biology professors, she found a position for the summer after sophomore year working in the research laboratory of a distinguished biochemist, Guido Guidotti. He turned her loose on a question that became her senior thesis: characterizing a protein that inexplicably churned up energy on the exterior of the cell.
“I learned an enormous amount from everyone in the Guidotti lab,” she says. “I got a sense of what it’s like to be a graduate student, and I realized it is very appealing to have that kind of freedom, exploration, and discovery. Guido also encouraged me to apply to the Stanford University Department of Biochemistry for grad school.”
ANIMAL DEVELOPMENT FROM THE OUTSIDE IN
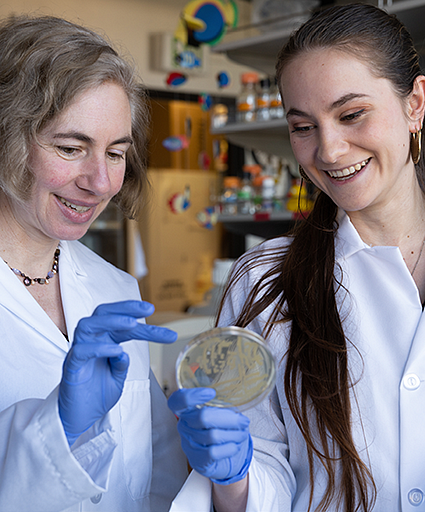

What happened next reveals how curious undergrads gradually transform into independent scientists who build on the (usually incremental) progress made by their predecessors. The process is similar to mastering a trade. Through hands-on immersion, they become proficient at delicate lab techniques, learn to construct experiments, and develop fluency in interpreting their data. As PhD candidates and postdoctoral fellows, they complete the circle by mentoring their own small teams of fledgling scientists.
During graduate school at Stanford, Guillemin studied with Mark Krasnow, a biochemist investigating the proteins that direct animal development. In her dissertation research on development of the fruit fly lung, Guillemin became intrigued by the external factors that direct animal development, as when flies living at high altitude grow lungs with more branches to deliver oxygen to their tissues. For her postdoctoral training she expanded these investigations to consider how bacteria influence animals in the context of infectious disease. She trained with the late Stanley Falkow at Stanford University School of Medicine, considered to be the founder of the field known as bacterial pathogenesis.
Throughout her postdoc, Guillemin knew she wanted to return to questions of animal development and explore the impacts of not just disease-causing but also beneficial microbes. She decided the best place to pursue her research was the UO. Just as many scientists must invent equipment needed to deepen their understanding of how something works, Guillemin knew the first step—raising fish to have zero microbes—was up to her.
The feat required a team effort. Guillemin worked with then-graduate student Jennifer Bates Atwood, BS ’02, PhD ’07 (biology), and had support from the UO’s fish experts, including colleagues of the late George Streisinger, the UO molecular biologist credited with establishing zebrafish as a powerful platform for genetic research.
“We were hypothesizing that bacteria were important for certain aspects of development, but we did not know what aspects those would be,” she says. “The really important moment was when we realized that we could reverse specific developmental delays by adding back certain bacteria.”
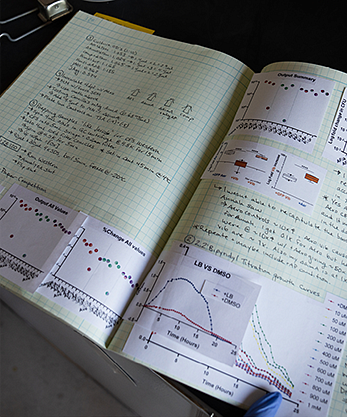
Bates Atwood’s and Guillemin’s findings also helped answer a longstanding puzzle—why our bodies don’t go into septic shock from our own bacteria—by revealing the peacekeeping effect of an intestinal enzyme that maintains friendly rapport between resident gut bacteria and their host.
“It was really exciting and rewarding because we were able to show that the host had very specific responses to the bacteria, and that these are all about coexisting with the bacteria,” she says. “For the first time, we saw how this could work at a molecular level.”
Today, Guillemin’s lab, which draws on the expertise of UO colleagues in fields ranging from physics to microbial ecology, is so productive that a list of discoveries emerging from it is beyond the scope of this article. For example, they realized a protein made by one type of bacteria reduces inflammation, which may lead to better therapeutics for Crohn’s disease and other intestinal disorders. They also found a bacterial protein that causes insulin-producing beta cells of the pancreas to multiply. Such cells are destroyed in people with Type 1 diabetes.
Her teams are also exploring strategies for derailing the bacteria known to cause ulcers and some types of stomach cancer. These ideas build on their discovery that this type of bacteria makes a protein that senses—and propels it toward—tissue that already is inflamed. This type of approach may eventually reduce the need for stronger antibiotics.
In addition, one of Guillemin’s research teams recently became the first from the Institute of Molecular Biology to tap into Lens on the Market, a UO program that helps scientists transition their research into commercial ventures. They are exploring the idea of offering screening services using zebrafish as the testing platform. “We can test many more microbes or microbial products using zebrafish than is possible using current animal models,” she says.
IT REALLY IS ALL META
In addition to collaborating with colleagues at the UO and around the world, Guillemin oversees teams of young scientists (including undergrads) and teaches microbiology. She also serves as founding director of the META Center for Host-Microbe Systems Biology, a one-of-a-kind federal research powerhouse established at the UO by the National Institutes of Health.
META stands for Microbial Ecology and Theory of Animals, and the acronym is apt. In science, “meta” designates work that encompasses multiple scales of investigation. Research at the META center seeks to understand the big picture of how microbes and animals interact, from molecular activity in cells to entire populations of animals, including humans. Scientists affiliated with the center grapple with big questions, such as how networks of microbial interactions connect all people and what makes microbes grow successfully inside us.
Guillemin says the spread of COVID-19 is a timely reminder that we live in an interconnected microbial world that spans national borders: “We can see the immense toll of the pandemic spread of this pathogen.” She points out that there may be other contagious microbes that confer some protection against COVID-19 by stimulating the right kind of immune response or even inhibiting viral replication.
“There is a real need for new strategies,” she says. “To achieve the goal of improving human health, we need to think like microbes.”
For a deep dive into how Guillemin is helping crystalize a new way of thinking about biomedical research, check out this article she coauthored in the April edition of Current Opinion in Microbiology.
Melody Ward Leslie, BA ’79 (humanities), is a staff writer for University Communications.
Photos by Dusty Whitaker, University Communications.
Fluoroscopy images by Travis Wiles, PhD.


