Scientists have known for decades that a certain class of enzymes are an important player in cell biology because they frequently mutate and become major drivers of cancer.
Biopharma companies are trying to develop drugs that target and inactivate these enzymes, known as phosphoinositide 3-kinase, or PI3K for short, because of their role in causing cancers in humans. But to do that, scientists need a detailed blueprint of the enzyme architecture, and UO biochemistry professor Scott Hansen is part of a group uncovering that diagram.
Hansen, an assistant professor in the UO Department of Chemistry and Biochemistry, said part of the challenge is that the molecules are complicated, with two large proteins that come together and form a complex.
“So it’s real challenging to figure out what is the structure and organization of this protein,” he said. “A prerequisite for developing a drug is learning how is the drug binding to the molecule. You need a blueprint of the protein to dock that drug on to.”
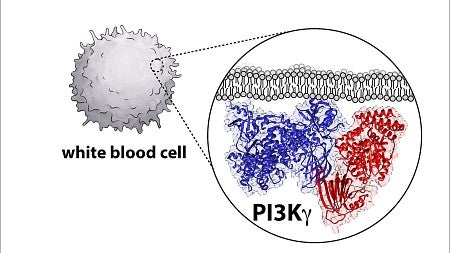
“This will definitely shape how people think about developing cancer therapeutics,” he said.
Hansen is one of 13 co-authors on the paper published in the journal Science Advances, along with colleagues at the universities of Victoria, British Columbia, Washington and Geneva, and Vrije Universiteit Brussel in Belgium.
What makes the paper unique is “higher resolution structural information and being able to measure biologically relevant interactions with other molecules in defining the sequence and priority of interactions,” Hansen said.
But he said much work remains to better understand how other molecules bind to the protein and modulate its function.
To learn the structure of the enzyme, the researchers used a technique called cryo-electron microscopy, which collects tens of thousands of images of single P13K enzymes in an ultra-thin layer of ice using an electron microscope. The images are compiled and averaged to create a single high-resolution image that reveals the structural features of the enzyme.
In this case, the structure of the enzyme was different than previously thought. The new blueprint will allow researchers to define how other molecules activate the protein.
“A lot of times we do research and we get clues about how proteins are organized or we look at structures of related molecules and extrapolate, but sometimes those assumptions are inaccurate and can lead a field down the wrong path,” Hansen said. “Having a higher resolution blueprint of the organization of molecules allows people to develop drugs specific to only that molecule.”
Hansen said researchers involved in the paper have spent 10 years working on the problem.
“That’s not uncommon for real complicated proteins,” he said.
Hansen said his lab at the UO and the lab of John Burke at the University of British Columbia conduct complementary research. Burke’s lab uses cryo-electron microscopy to get high-resolution images of molecules, and Hansen’s lab is able to “make more dynamic measures that allow us to measure the interactions of this enzyme and other factors critical to activation.”
He said it’s exciting to collaborate with other scientists who have expertise in areas that he does not.
“More and more, for us to accomplish our research ambitions we can’t just rely on the skills in our own lab,” he said. “Problems are getting too complicated and you have to approach them from different angles and integrate a lot of different expertise.”
—By Tim Christie, University Communications